Interact with mrgsimsds objects
Usage
# S3 method for class 'mrgsimsds'
dim(x)
# S3 method for class 'mrgsimsds'
head(x, n = 6L, ...)
# S3 method for class 'mrgsimsds'
tail(x, n = 6L, ...)
# S3 method for class 'mrgsimsds'
names(x)
# S3 method for class 'mrgsimsds'
plot(
x,
y = NULL,
...,
nid = 16,
batch_size = 20000,
logy = FALSE,
.dots = list()
)Arguments
- x
an mrgsimsds object, output from
mrgsim_ds()oras_mrgsim_ds().- n
number of rows to return.
- ...
arguments to be passed to or from other methods.
- y
a formula for plotting simulated data; if not provided, all columns will be plotted.
- nid
number of subjects to plot.
- batch_size
size of batch when reading data for plot method.
- logy
if
TRUE, plot data with log y-axis.- .dots
a list of items to pass to
mrgsolve::plot_sims().
Details
head() and tail() only look at the first and last file in the data
set, respectively, when simulations are stored across multiple files. It is
possible this won't correspond to the first and last chunks rows of data
you will see when collecting the data via dplyr::collect().
Examples
mod <- house_ds(end = 24)
mod <- omat(mod, diag(0.04, 4))
data <- ev_expand(amt = c(100, 300), ID = 1:20)
set.seed(10203)
out <- mrgsim_ds(mod, data = data)
dim(out)
#> [1] 3920 7
head(out)
#> # A tibble: 6 × 7
#> ID time GUT CENT RESP DV CP
#> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl>
#> 1 1 0 0 0 70.9 0 0
#> 2 1 0 100 0 70.9 0 0
#> 3 1 0.25 68.8 30.9 68.9 1.94 1.94
#> 4 1 0.5 47.3 51.7 65.1 3.24 3.24
#> 5 1 0.75 32.6 65.5 61.1 4.10 4.10
#> 6 1 1 22.4 74.5 57.6 4.66 4.66
tail(out)
#> # A tibble: 6 × 7
#> ID time GUT CENT RESP DV CP
#> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl>
#> 1 40 22.8 1.81e-11 124. 39.1 5.57 5.57
#> 2 40 23 1.28e-11 123. 39.2 5.52 5.52
#> 3 40 23.2 9.11e-12 122. 39.4 5.46 5.46
#> 4 40 23.5 6.44e-12 121. 39.5 5.41 5.41
#> 5 40 23.8 4.56e-12 119. 39.7 5.36 5.36
#> 6 40 24 3.23e-12 118. 39.8 5.30 5.30
nrow(out)
#> [1] 3920
ncol(out)
#> [1] 7
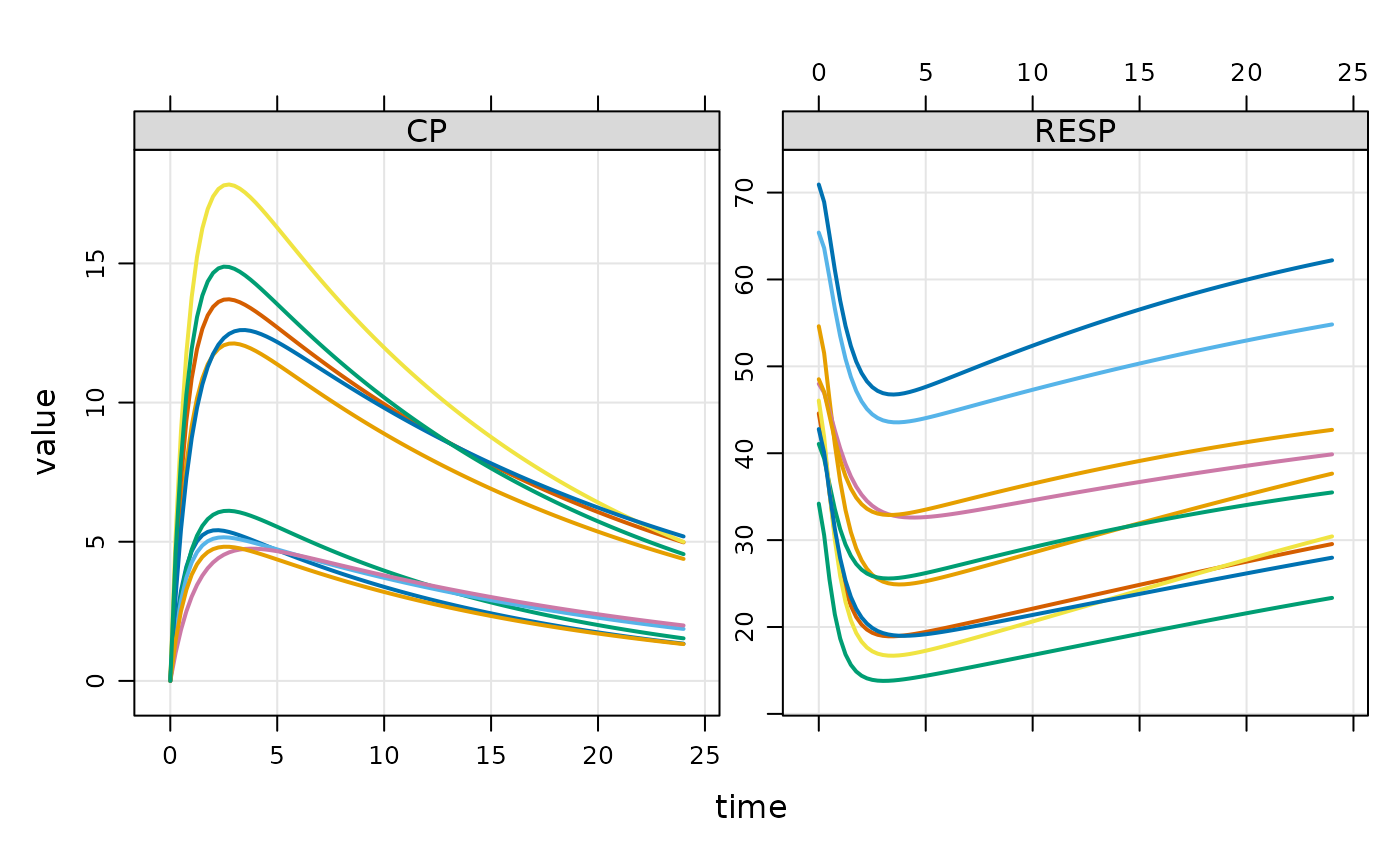
plot(out, ~ CP + RESP, nid = 10)