mrgsim.ds provides an Apache Arrow-backed simulation output object for mrgsolve, greatly reducing the memory footprint of large simulations and providing a high-performance pipeline for summarizing huge simulation outputs. The arrow-based simulation output objects in R claim ownership of their files on disk. Those files are automatically removed when the owning object goes out of scope and becomes subject to the R garbage collector. While “anonymous”, parquet-formatted files hold the data in tempdir() as you are working in R, functions are provided to move this data to more permanent locations for later use.
Installation
You can install the development version of mrgsim.ds from r-universe with:
# Install 'mrgsim.ds' in R:
install.packages('mrgsim.ds', repos = c('https://kylebaron.r-universe.dev', 'https://cloud.r-project.org'))Example
We will illustrate mrgsim.ds by doing a simulation.
library(mrgsim.ds)
library(dplyr)
mod <- modlib_ds("popex", end = 240, outvars = "IPRED,CL")
data <- expand.ev(amt = 100, ii = 24, total = 6, ID = 1:3000)mrgsim.ds provides a new mrgsim() variant - mrgsim_ds(). The name implies we are tapping into Apache Arrow Dataset functionality. The simulation below carries 1,446,000 rows.
out <- mrgsim_ds(mod, data)
out
. Model: popex
. Dim : 1.4M x 4
. Files: 1 [11.9 Mb]
. Owner: yes
. ID time CL IPRED
. 1: 1 0.0 0.6601045 0.000000
. 2: 1 0.0 0.6601045 0.000000
. 3: 1 0.5 0.6601045 1.756330
. 4: 1 1.0 0.6601045 2.947337
. 5: 1 1.5 0.6601045 3.744798
. 6: 1 2.0 0.6601045 4.268478
. 7: 1 2.5 0.6601045 4.601877
. 8: 1 3.0 0.6601045 4.803204Very lightweight simulation output object
The output object doesn’t actually carry these 1.4M rows of simulated data. Rather it stores a pointer to the data in parquet files on your disk.
This means there is almost nothing inside the object itself
What if we did the same simulation with regular mrgsim()?
The mrgsim.ds object is very light weight despite tracking the same data.
Handles like regular mrgsim output
But, we can do a lot of the typical things we would with any mrgsim() output object.
plot(out, nid = 12)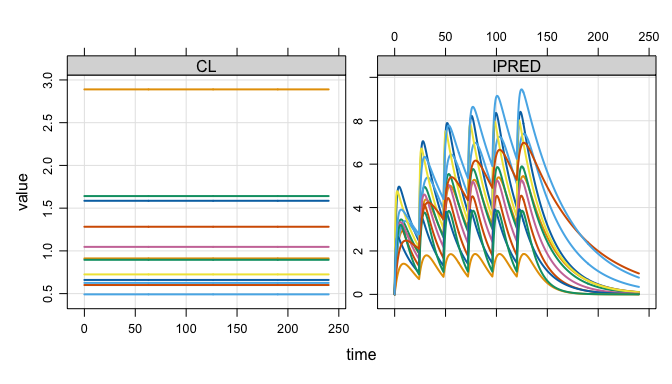
head(out)
. # A tibble: 6 × 4
. ID time CL IPRED
. <dbl> <dbl> <dbl> <dbl>
. 1 1 0 0.660 0
. 2 1 0 0.660 0
. 3 1 0.5 0.660 1.76
. 4 1 1 0.660 2.95
. 5 1 1.5 0.660 3.74
. 6 1 2 0.660 4.27
tail(out)
. # A tibble: 6 × 4
. ID time CL IPRED
. <dbl> <dbl> <dbl> <dbl>
. 1 3000 238. 0.779 0.119
. 2 3000 238 0.779 0.117
. 3 3000 238. 0.779 0.115
. 4 3000 239 0.779 0.113
. 5 3000 240. 0.779 0.111
. 6 3000 240 0.779 0.109
dim(out)
. [1] 1446000 4This includes coercing to different types of objects. We can get the usual R data frames
as_tibble(out)
. # A tibble: 1,446,000 × 4
. ID time CL IPRED
. <dbl> <dbl> <dbl> <dbl>
. 1 1 0 0.660 0
. 2 1 0 0.660 0
. 3 1 0.5 0.660 1.76
. 4 1 1 0.660 2.95
. 5 1 1.5 0.660 3.74
. 6 1 2 0.660 4.27
. 7 1 2.5 0.660 4.60
. 8 1 3 0.660 4.80
. 9 1 3.5 0.660 4.91
. 10 1 4 0.660 4.96
. # ℹ 1,445,990 more rowsOr stay in the arrow ecosystem
as_arrow_ds(out)
. FileSystemDataset with 1 Parquet file
. 4 columns
. ID: double
. time: double
. CL: double
. IPRED: double
.
. See $metadata for additional Schema metadataOr try your hand at duckdb
as_duckdb_ds(out)
. # Source: table<arrow_001> [?? x 4]
. # Database: DuckDB 1.4.3 [kyleb@Darwin 24.6.0:R 4.5.2/:memory:]
. ID time CL IPRED
. <dbl> <dbl> <dbl> <dbl>
. 1 1 0 0.660 0
. 2 1 0 0.660 0
. 3 1 0.5 0.660 1.76
. 4 1 1 0.660 2.95
. 5 1 1.5 0.660 3.74
. 6 1 2 0.660 4.27
. 7 1 2.5 0.660 4.60
. 8 1 3 0.660 4.80
. 9 1 3.5 0.660 4.91
. 10 1 4 0.660 4.96
. # ℹ more rowsTidyverse-friendly
We’ve integrated into the dplyr ecosystem as well, allowing you to filter(), group_by(), mutate(), select(), summarise(), rename(), or arrange() your way directly into a pipeline to summarize your simulations using the power of Apache Arrow.
Good for large simulations
This workflow is particularly useful when running replicate simulations in parallel, with large outputs
library(future.apply, quietly = TRUE)
plan(multisession, workers = 5L)
out2 <- future_lapply(1:10, \(x) { mrgsim_ds(mod, data) }, future.seed = TRUE)
out2 <- reduce_ds(out2)Now there are 10x the number of rows (14.5M), but little change in object size.
out2
. Model: popex
. Dim : 14.5M x 4
. Files: 10 [119.2 Mb]
. Owner: yes
. ID time CL IPRED
. 1: 1 0.0 0.7453202 0.0000000
. 2: 1 0.0 0.7453202 0.0000000
. 3: 1 0.5 0.7453202 0.2004743
. 4: 1 1.0 0.7453202 0.3821993
. 5: 1 1.5 0.7453202 0.5467789
. 6: 1 2.0 0.7453202 0.6956808
. 7: 1 2.5 0.7453202 0.8302481
. 8: 1 3.0 0.7453202 0.9517104Files on disk are automagically managed
All arrow files are stored in the tempdir() in parquet format
list_temp()
. 11 files [131.1 Mb]
. - mrgsims-ds-6ece3b5bb339.parquet
. - mrgsims-ds-6f0c360fd6d3.parquet
. ...
. - mrgsims-ds-6f10225ef974.parquet
. - mrgsims-ds-6f1094b404d.parquetThis directory is eventually removed when the R session ends. Tools are provided to manage the space.
retain_temp(out2)
. Discarding 1 files.
list_temp()
. 10 files [119.2 Mb]
. - mrgsims-ds-6f0c360fd6d3.parquet
. - mrgsims-ds-6f0c49e0d25f.parquet
. ...
. - mrgsims-ds-6f10225ef974.parquet
. - mrgsims-ds-6f1094b404d.parquetWe also put a finalizer on each object so that, when it goes out of scope, the files are automatically cleaned up.
First, run a bunch of simulations.
plan(multisession, workers = 5L)
out1 <- mrgsim_ds(mod, data)
rename_ds(out1, "out1")
out2 <- future_lapply(1:10, \(x) { mrgsim_ds(mod, data) }, future.seed = TRUE)
out2 <- reduce_ds(out2)
rename_ds(out2, "out2")
out3 <- mrgsim_ds(mod, data)
rename_ds(out3, "out3")There are 12 files holding simulation outputs.
list_temp()
. 12 files [143 Mb]
. - mrgsims-ds-out1-0001.parquet
. - mrgsims-ds-out2-0001.parquet
. ...
. - mrgsims-ds-out2-0010.parquet
. - mrgsims-ds-out3-0001.parquetNow, remove one of the objects containing 10 files.
rm(out2)As soon as the garbage collector is called, the leftover files are cleaned up.
gc()
. used (Mb) gc trigger (Mb) limit (Mb) max used (Mb)
. Ncells 1946964 104.0 3643540 194.6 NA 3271222 174.8
. Vcells 15237389 116.3 29085557 222.0 16384 27013458 206.1
list_temp()
. 2 files [23.8 Mb]
. - mrgsims-ds-out1-0001.parquet
. - mrgsims-ds-out3-0001.parquetOwnership
This setup is only possible if one object owns the files on disk and mrgsim.ds tracks this.
If I make a copy of a simulation object, the old object no longer owns the files.
out4 <- copy_ds(out1, own = TRUE)
check_ownership(out1)
. [1] FALSE
check_ownership(out4)
. [1] TRUEI can always take ownership back.
If this is so great, why not make it the default for mrgsolve?
There is a cost to all of this. For small to mid-size simulations, you might see a small slowdown with mrgsim_ds(); it definitely won’t be faster than mrgsim() … even with the super-quick arrow ecosystem. This workflow is really for large simulation volumes where you are happy to pay the cost of writing outputs to file and then streaming them back in to summarize.